Mitchell O’Leary1, Avi S. Baskin, MD MPhil1
Department of Urology, UC Irvine Health 101 The City Drive South, Pavilion III, Building 29, Orange, CA 92868
Corresponding Author: Avi S. Baskin, abaskin@hs.uci.edu
Keywords: small renal mass; renal cell carcinoma; molecular biomarkers; risk stratification; genomic profiling
ABSTRACT
BACKGROUND AND OBJECTIVE: Small renal masses (SRMs), defined as enhancing renal lesions ≤4 cm (clinical stage T1a), represent a biologically heterogeneous group of tumors. While many SRMs are benign or indolent, a subset harbor aggressive potential, and current imaging modalities cannot reliably distinguish these subgroups. This narrative review summarizes the current evidence for molecular and genomic biomarkers in SRM risk stratification and evaluates their potential clinical role.
METHODS: A structured search of PubMed/MEDLINE from January 1, 2000 through January 1, 2026 was performed using terms related to small renal masses, molecular and genomic biomarkers, and prognostic outcomes. Reference lists of relevant studies and clinical guidelines were also reviewed.
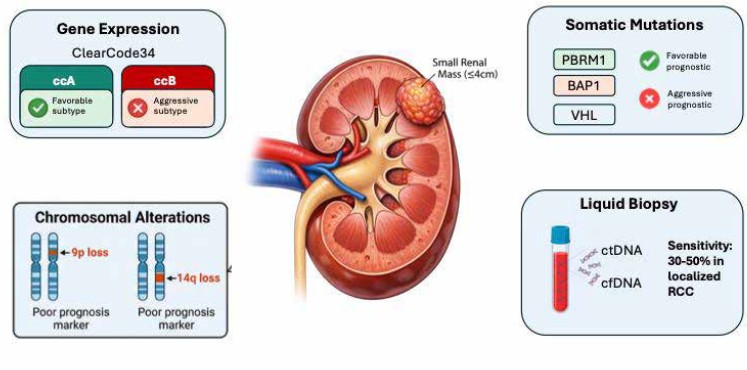
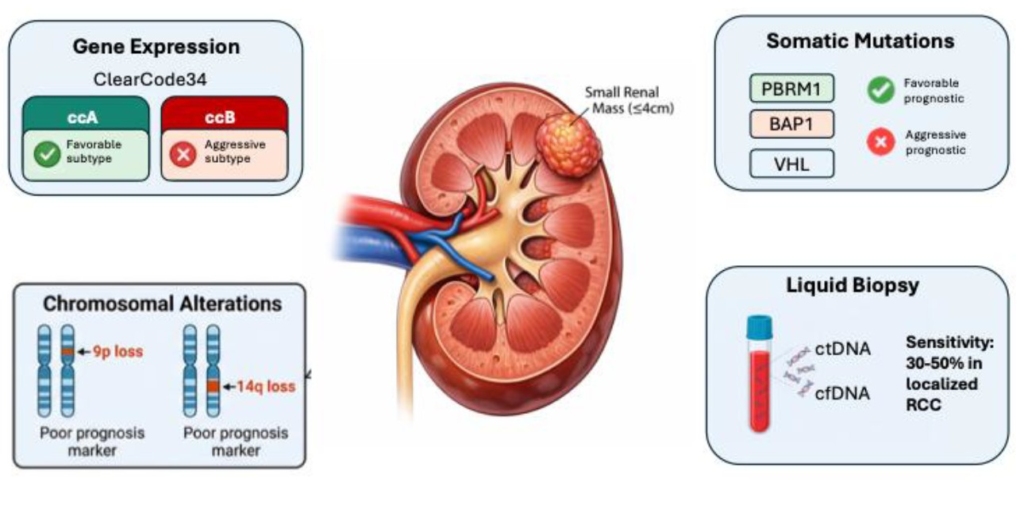
RESULTS: Gene expression signatures, including ClearCode34 and a 16-gene recurrence assay, can distinguish indolent from aggressive clear cell renal cell carcinoma and are compatible with formalin-fixed biopsy tissue. Somatic alterations in genes such as VHL, PBRM1, BAP1, and SETD2 define biologically distinct tumor subtypes, with BAP1 loss detectable by immunohistochemistry on renal mass biopsy specimens. Chromosomal alterations, including 9p loss, correlate with higher-grade disease and metastatic potential. Emerging liquid biopsy approaches demonstrate promise but remain incompletely validated in SRM populations.
CONCLUSIONS: Although molecular and genomic biomarkers have substantially advanced understanding of renal cell carcinoma biology, none are currently incorporated into routine clinical management of small renal masses. Most candidate biomarkers have been developed in nephrectomy cohorts and lack prospective validation in renal mass biopsy specimens or active surveillance populations. Future studies integrating molecular biomarkers with clinical and radiographic data in SRM-specific cohorts are needed before these tools can meaningfully inform patient care.
Introduction
Renal cell carcinoma (RCC) accounts for over 80,000 new diagnoses and 14,500 deaths in the United States annually.1 Given the increased use of cross sectional imaging, renal tumors are increasingly detected at an earlier stage, with small renal masses (SRMs), defined as incidentally detected contrast enhancing renal lesions ≤4 cm (clinical stage cT1a) comprising 50-65% of newly diagnosed cases.23 This stage migration, while oncologically favorable, has created significant challenges in individualized risk stratification. Notably, despite increased early detection, improvements in overall and disease-specific survival have not been proportionate, suggesting that earlier detection alone is insufficient and that improved biological characterization is needed to guide individualized management decisions.4,5
The biological spectrum of SRMs is markedly heterogeneous.6 The 2022 World Health Organization classification now recognizes over 20 RCC subtypes, including a new category of molecularly defined entities, reflecting the increasingly appreciated molecular diversity underlying morphologically similar renal tumors.7 Approximately 20-30% of biopsied SRMs are benign (most commonly oncocytoma and angiomyolipoma) while the majority are malignant, predominantly clear cell RCC (ccRCC), papillary or chromophobe histology.8,9 Among malignant SRMs, most pursue an indolent clinical course amenable to active surveillance, while a clinically significant minority harbor high-risk features including high nuclear grade, aggressive histologic subtypes, or adverse molecular profiles.5,10,11 Current standard cross-sectional imaging cannot reliably differentiate indolent from aggressive SRMs, contributing to overtreatment in a predominantly low-risk disease population.4,9
Renal mass biopsy (RMB) has emerged as a valuable adjunct for risk stratification providing histologic diagnosis in 80-90% of cases with a low complication profile.12,13 However, histology alone provides limited prognostic information beyond tumor grade and subtype. The integration of molecular and genomic biomarkers from biopsy specimens—including gene expression signatures, somatic mutation profiling, and epigenetic markers—represents a compelling strategy to augment risk stratification and guide individualized management. Current management recommendations from the American Urological Association and the European Association of Urology endorse RMB as part of the evaluation of SRMs but acknowledge the absence of validated molecular adjuncts.
From a clinical perspective, molecular biomarkers in SRMs may serve several distinct purposes. First, biomarkers may help distinguish benign from malignant renal masses, potentially reducing overtreatment of benign lesions such as oncocytoma. Second, biomarkers may aid in histologic subtype classification when RMB specimens are limited or indeterminate. Third, molecular features may help identify tumors likely to demonstrate rapid growth during active surveillance. Finally, biomarkers may assist in predicting aggressive biology or metastatic potential. Importantly, biomarkers developed for these different clinical endpoints are not interchangeable, and understanding the specific clinical context in which a biomarker is intended to operate is essential when evaluating the current literature.
In this narrative review, we synthesize the evidence for molecular and genomic biomarkers in risk stratification of SRMs, organized by biomarker category: gene expression signatures, somatic mutations, chromosomal alterations, and liquid biopsy. We additionally provide an expert assessment of the clinical implications of these data and directions for future investigation.
Methods
Literature Search
A structured literature search of PubMed/MEDLINE (National Library of Medicine) was performed from 1/1/2000 through 1/1/2026, reflecting the era in which molecular profiling technologies applicable to renal tumor characterization have become available. The search strategy combined Medical Subject Headings (MeSH) terms and free-text keywords organized into three concept blocks: (1) the population (small renal mass, renal mass, T1a, localized renal cell carcinoma), (2) the index biomarker (molecular biomarker, gene expression, somatic mutation, liquid biopsy, epigenetic, microRNA, and specific gene names including VHL, PBRM1, BAP1, SETD2, and ClearCode34), and (3) the outcome context (risk stratification, prognosis, active surveillance, metastasis-free survival, cancer-specific survival). Concept blocks were combined with the Boolean AND operator and limited to human studies in the English language. The complete search string is provided in Supplementary Table S1.
The search was supplemented by manual review of reference lists from all included studies, relevant narrative reviews, and current clinical practice guidelines from the American Urological Association (AUA) and European Association of Urology (EAU) to identify additional eligible studies.
Study Selection
Studies were included if they: (1) evaluated at least one molecular or genomic biomarker (gene expression signature, somatic mutation, chromosomal alteration, liquid biopsy analyte) in renal tumors ≤4 cm or clinical T1a; (2) reported a prognostic, diagnostic, or predictive endpoint; and (3) were published as full-text articles in peer-reviewed journals. Studies including broader localized RCC cohorts were included if SRM-relevant data were reported or extractable. Case reports, conference abstracts, preclinical studies, and studies with fewer than 10 patients were excluded. Studies focused exclusively on advanced or metastatic RCC without SRM-applicable data were excluded.
Data Synthesis
Results are presented as a narrative synthesis organized by biomarker category: gene expression signatures, somatic mutation profiling, chromosomal copy number alterations, and liquid biopsy markers. Epigenetic and microRNA-based biomarkers are discussed briefly in the context of future directions but were not included as primary evidence categories, as the available literature is largely confined to metastatic or nephrectomy cohorts without SRM-specific data.. Within each category, the available evidence is summarized with emphasis on analytical validation status, clinical endpoints, and applicability to biopsy-derived specimens. Artificial Intelligence tools (Claude, Anthropic, ChatGPT, OpenAI) were used to assist with figure generation and proofreading. All outputs were verified by authors. The literature search was executed, and study selection was performed by the authors.
Results
The structured PubMed search identified 1,277 records. After removal of duplicates, title and abstract screening excluded 1,180 records; full-text review of 97 articles resulted in 38 studies meeting inclusion criteria, forming the basis of this review, spanning four biomarker categories: gene expression signatures, somatic mutation profiling, chromosomal copy number alterations, and liquid biopsy markers.
Gene Expression Signatures
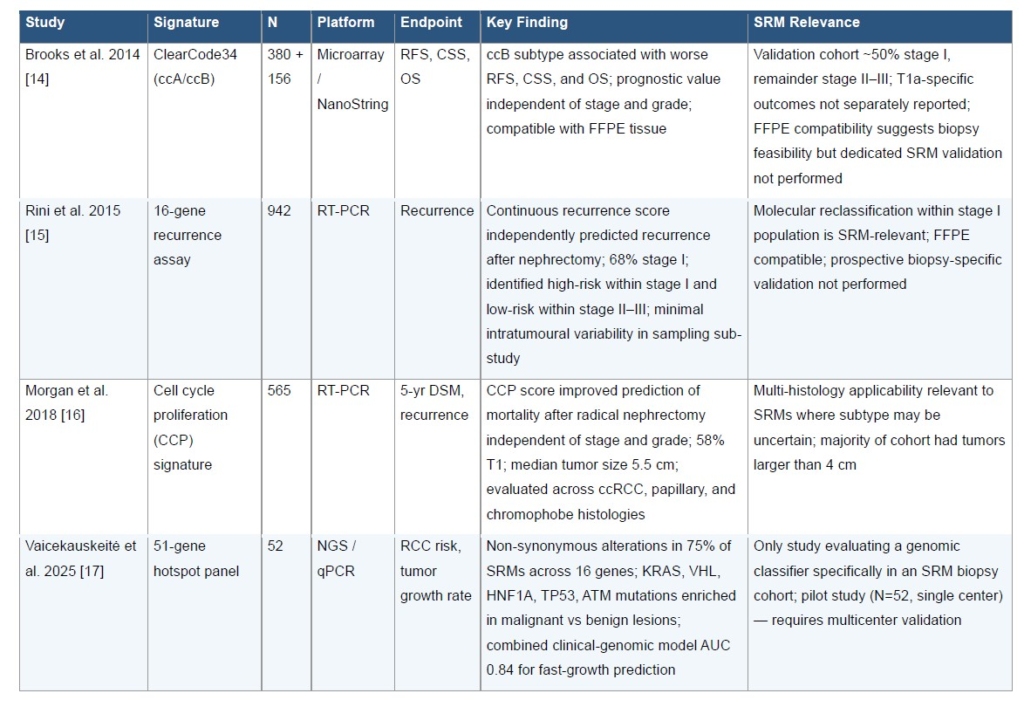
Gene expression profiling has yielded several clinically relevant signatures applicable to SRM stratification (Table 1). ClearCode34, a 34-gene expression signature derived from The Cancer Genome Atlas (TCGA) ccRCC data, classifies tumors into two molecular subtypes: ccA (associated with favorable prognosis) and ccB (associated with aggressive biology). In the index study, the classifier was applied to 380 non-metastatic ccRCC samples from TCGA and validated in 156 formalin-fixed clinical samples, demonstrating significant associations with recurrence-free survival, cancer-specific survival, and overall survival.14 It is important to note that the validation cohort included tumors across the full spectrum of localized disease, approximately 50% stage I with the remainder stage II-III. While cT1a-specific outcomes were not separately reported, the signature’s prognostic value after adjustment for tumor size and its compatibility with FFPE tissue suggest potential applicability to biopsy-based SRM assessment.
Rini et al. developed and validated a 16-gene recurrence assay for localized ccRCC in a multi-institutional study of 942 patients with stage I–III disease.15 The assay, based on reverse transcription-polymerase chain reaction quantification of genes involved in angiogenesis, immune response, and cell proliferation, generated a continuous recurrence score that independently predicted recurrence after nephrectomy. Notably, 68% of the development cohort had stage I disease, and the recurrence score identified high-risk patients within stage I and low-risk patients within stage II–III, demonstrating molecular reclassification beyond conventional staging. The assay was designed for FFPE surgical specimens and showed minimal intratumoral variability in a sampling sub-study, though prospective validation on core needle biopsy material has not been performed.
Morgan et al. developed a multigene cell cycle proliferation (CCP) signature and demonstrated improved prediction of 5-year disease-specific mortality after radical nephrectomy in a study of 565 patients across two independent cohorts (Massachusetts General Hospital and University of Michigan).16 Unlike other gene expression classifiers restricted to ccRCC, the CCP score was evaluated across clear cell, papillary, and chromophobe histologies, with 58% of patients having T1 disease. The CCP score provided prognostic information independent of stage and grade, suggesting potential utility in the SRM population where histologic subtype may be uncertain prior to definitive intervention.
Somatic Mutation Profiling
A 2025 study by Vaicekauskeitė et al. evaluated the genomic landscape of SRMs specifically, sequencing 51 cancer hotspot genes in 52 patients.17 Non-synonymous alterations were identified in 75% of SRMs, with KRAS, VHL, HNF1A, TP53, and ATM mutations predominantly detected in malignant versus benign lesions. A combined clinical-genomic model achieved an area under the curve (AUC) of 0.84 for prediction of rapid tumor growth, supporting the feasibility of biopsy-based hotspot panel testing in SRM management. Notably, this is the only study in the current literature to evaluate a genomic classifier specifically in an SRM biopsy cohort, though the small sample size and single-center design warrant validation in larger multicenter studies.
The somatic mutation landscape of ccRCC has been extensively characterized in surgical nephrectomy cohorts spanning all stages of localized and advanced disease. While SRM-specific mutational profiling data remain limited, the biological and prognostic significance of these mutations provides the rationale for their potential application to SRM risk stratification. (Table 2)
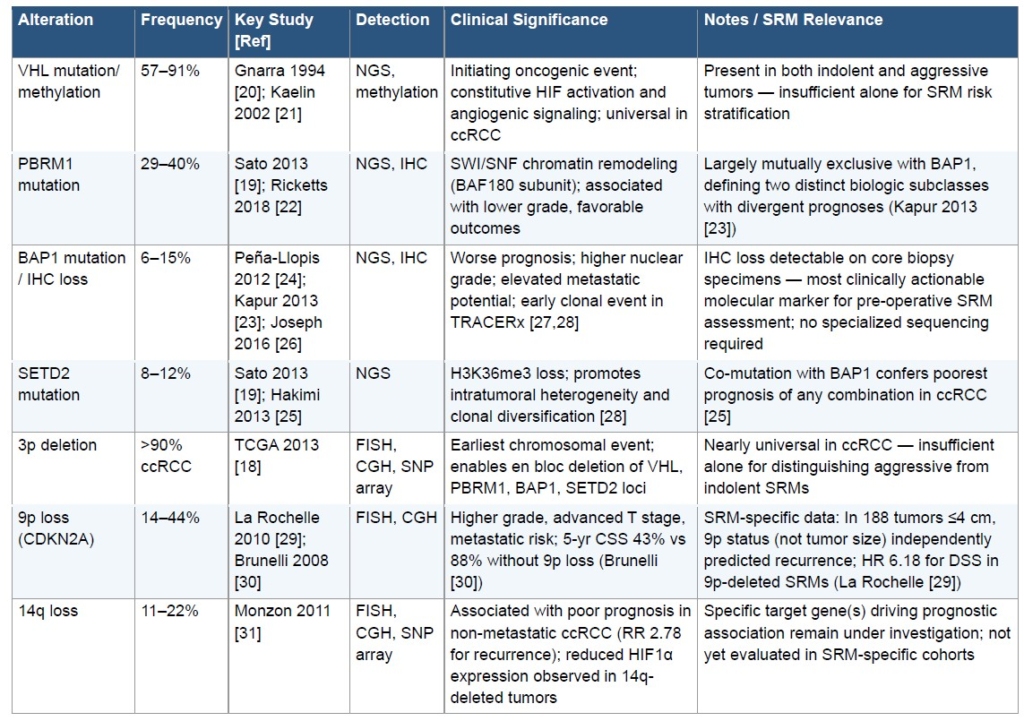
The genomic landscape of ccRCC is characterized by recurrent driver mutations concentrated on chromosome 3p21.18,19 VHL mutations or promoter methylation are present in 57–91% of sporadic ccRCCs and represent the initiating oncogenic event, driving constitutive HIF activation and downstream angiogenic signaling.20,21 However, VHL loss is essentially universal in ccRCC—including biologically indolent tumors—and therefore lacks discriminatory value for SRM risk stratification when used in isolation.
PBRM1 mutations, present in approximately 29–40% of ccRCCs, encode the BAF180 subunit of the SWI/SNF chromatin remodeling complex.19,22 PBRM1-mutated tumors tend toward lower grade and stage and are associated with more favorable outcomes. Critically, PBRM1 and BAP1 mutations are largely mutually exclusive in ccRCC, defining two distinct biologic subclasses with divergent prognoses. 23
BAP1 mutations, occurring in 6–15% of ccRCCs, encode a nuclear deubiquitinase regulating chromatin compaction and DNA damage repair.24 BAP1-mutated ccRCCs demonstrate consistently worse outcomes across multiple independent cohorts, with higher nuclear grade, elevated metastatic potential, and inferior cancer-specific survival. 23,25 Loss of BAP1 protein expression is detectable by immunohistochemistry (IHC) on core biopsy specimens, providing a clinically actionable prognostic marker applicable to pre-operative SRM assessment.23,26 The landmark TRACERx Renal study demonstrated that BAP1 loss is an early clonal event correlated with metastatic evolutionary trajectories, supporting its association with aggressive biology.27,28
SETD2 mutations, encoding the histone methyltransferase responsible for H3K36 trimethylation, occur in 8–12% of ccRCCs.19,25 SETD2 loss promotes intratumoral heterogeneity and clonal diversification.28 Co-occurrence of BAP1 and SETD2 mutations confers the poorest prognosis of any mutational combination in ccRCC, reflecting convergent disruption of chromatin remodeling and DNA damage surveillance pathways.25
The hierarchical clonal evolution model of ccRCC — in which VHL loss represents the trunk mutation and subsequent drivers (PBRM1, BAP1, SETD2, KDM5C) define branch-level heterogeneity — has direct implications for biopsy-based SRM assessment.18,27,28 The TRACERx Renal study demonstrated that driver mutations such as BAP1 loss can be regionally distributed within a single tumor, implying that single-core biopsies may underestimate aggressive subclonal populations.27,28 Optimized multicore sampling strategies targeting the periphery and solid tumor components may be necessary to achieve reliable molecular classification from biopsy specimens.
Chromosomal alterations
Chromosomal copy number alterations provide an additional layer of prognostic information applicable to SRM risk stratification (Table 2). Loss of chromosome 3p, occurring in >90% of ccRCCs, is the defining early event enabling deletion of VHL, PBRM1, BAP1, and SETD2 en bloc.18 However, as with VHL mutation, 3p loss is essentially universal in ccRCC and lacks specificity for distinguishing aggressive from indolent SRMs.
Loss of chromosome 9p is among the most SRM-relevant chromosomal biomarkers identified to date. La Rochelle et al. analyzed 703 ccRCC tumors, including a subgroup analysis of 188 patients with ≤4 cm tumors, and found that 9p deletion independently predicted worse disease-specific and recurrence-free survival.29 Critically, among small localized tumors, 9p status, but not tumor size, was independently associated with disease recurrence, raising the possibility of using preoperative FISH analysis on biopsy specimens to identify aggressive biology within otherwise indolent-appearing SRMs.29 These findings were consistent with earlier work by Brunelli et al., who demonstrated that 9p loss was an independent prognostic factor with 5-year cancer-specific survival of 43% versus 88% in patients without 9p loss.30
Chromosome 14q loss, occurring in approximately 11–22% of ccRCCs, has been associated with poor prognosis in non-metastatic disease, with an increased risk of recurrence (RR 2.78) and decreased overall survival.31 The HIF1A gene resides on 14q, and reduced HIF1α expression has been observed in 14q-deleted tumors, though the specific target gene(s) driving this association remain under investigation. Whether 14q status adds incremental value to 9p assessment in the SRM population has not been evaluated.
Liquid Biopsy
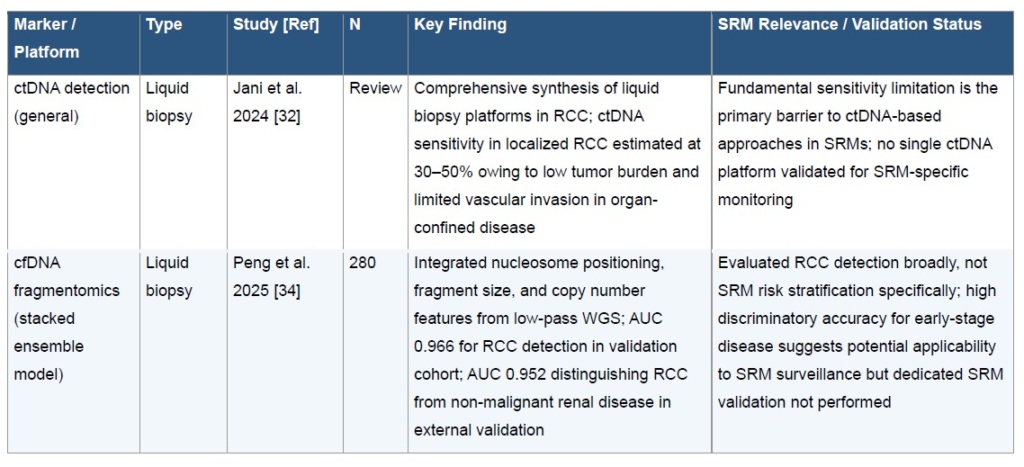
Liquid biopsy represents an emerging non-invasive strategy with theoretical applicability to SRM monitoring, though the current evidence base is limited. (Table 3) The majority of circulating tumor DNA (ctDNA) data in RCC has been generated in the metastatic setting, where tumor shedding is substantially higher than in localized disease. In localized RCC and SRMs specifically, ctDNA detection sensitivity is estimated at only 30–50%, reflecting the low tumor burden and limited vascular invasion characteristic of small, organ-confined tumors.32,33 This fundamental sensitivity limitation represents the primary barrier to clinical translation of ctDNA-based approaches in the SRM population.
Cell-free DNA (cfDNA) fragmentomics, which analyzes nucleosome positioning patterns, fragment size distributions, and copy number variations using machine learning, may offer a path to overcoming this sensitivity gap. Peng et al. developed a stacked ensemble model integrating three cfDNA features from low-pass whole-genome sequencing in 280 participants, achieving an AUC of 0.966 for RCC detection and 0.952 for distinguishing RCC from non-malignant renal disease in external validation.34 While this study evaluated RCC detection broadly and did not include an SRM-specific subgroup analysis, the high discriminatory accuracy for early-stage disease suggests potential applicability to the SRM surveillance setting, where differentiating indolent from progressive disease without repeat biopsy remains an unmet clinical need.
A 2024 synthesis of liquid biopsy platforms in RCC by Jani et al. concluded that no single platform has been validated for SRM-specific monitoring, and that prospective studies evaluating serial liquid biopsy in the active surveillance SRM population represent a critical research gap.32
Discussion
This narrative review synthesizes the current evidence for molecular and genomic biomarkers across four categories: gene expression signatures, somatic mutation profiling, chromosomal alterations, and liquid biopsy. While advances in genomic sequencing and transcriptomic profiling have substantially improved understanding of RCC biology, translating these findings into actionable tools for SRM management remains a significant challenge.
Most studies evaluating genomic alterations and transcriptomic classifiers have been conducted in nephrectomy cohorts that include tumors across a wide size spectrum, many of which are substantially larger than typical SRMs.5,14 Consequently, the applicability of these findings to incidentally detected tumors ≤4 cm remains uncertain. SRMs exhibit a broad biological spectrum, with a substantial proportion representing benign lesions or indolent malignancies that may be safely managed with active surveillance.5 As a result, biomarkers developed in cohorts enriched for larger or more advanced tumors may not accurately reflect the biological heterogeneity present within the SRM population. Prospective validation in cohorts specifically composed of SRMs is therefore necessary before molecular biomarkers can be integrated into routine clinical decision-making.
A major challenge in applying molecular biomarkers to SRM management is the issue of intratumoral heterogeneity and the limitations inherent to RMB. Multiregion sequencing studies have demonstrated that individual RCC tumors can contain substantial spatial genomic heterogeneity, with distinct tumor regions harboring different mutational profiles.35 Work from the TRACERx Renal Consortium further demonstrated that while some genomic alterations represent early truncal events in tumor evolution, many potentially prognostic alterations occur later in tumor evolution and may therefore be regionally restricted.27,28 RMB samples only a small portion of the tumor and these spatial differences raise concern that a single biopsy specimen may not capture the complete molecular landscape of the tumor. Identifying biomarkers that represent early truncal events and are therefore consistently detectable across tumor regions may be critical for successful clinical implementation.
From a clinical perspective, the most relevant application of molecular biomarkers in SRMs is their potential to improve the decisions clinicians and patients must make at presentation, rather than to predict long-term oncologic outcomes in isolation. In particular, biomarkers that help distinguish benign from malignant tumors, or indolent from potentially aggressive lesions, could reduce overtreatment in a disease space where many masses are either benign or biologically low risk.8,10,11 This is especially important in the setting of active surveillance, where current decision-making relies heavily on tumor size, growth kinetics, and biopsy histology, each of which has important limitations. 9-13 Molecular biomarkers may therefore be most clinically useful not as stand-alone tests, but as adjuncts to existing clinical, radiographic, and pathologic data. Early SRM-specific studies, including recent genomic analyses linking certain mutational profiles with more rapid tumor growth, support the possibility that integrated molecular-clinical models could eventually refine surveillance selection and identify the subset of patients most likely to benefit from intervention.17
Future progress in applying molecular biomarkers to SRMs will likely depend on improved tissue acquisition strategies and prospective validation within SRM-specific cohorts. RMB has become increasingly incorporated into clinical decision-making, yet current biopsy techniques often yield limited tissue and may inadequately capture tumor heterogeneity. 12,13 More systematic biopsy approaches, including obtaining multiple cores from spatially distinct tumor regions, may improve molecular characterization and better reflect the underlying tumor biology.
At the same time, the inherent limitations of tissue sampling highlight the potential value of noninvasive biomarker approaches. Liquid biopsy technologies capable of detecting ctDNA, circulating tumor cells, or tumor-derived RNA in blood or urine offer an attractive complementary strategy for characterizing tumor biology and monitoring disease over time. Although current studies suggest that ctDNA levels are often low in localized RCC, advances in assay sensitivity and detection methods may improve the feasibility of these approaches in SRMs.32,36
Ultimately, the most promising strategy may involve integrating molecular data from tissue, liquid biopsy platforms, and clinical variables into multimodal predictive models. Such approaches have already begun to transform risk stratification in other cancers and may similarly enable more precise and individualized management of patients with SRMs. Another promising avenue is the integration of imaging characteristics with molecular data through radiogenomic approaches, which aim to identify imaging signatures that correlate with underlying tumor genomic alterations.37 Epigenetic and microRNA-based biomarkers have also demonstrated biologic and prognostic relevance in RCC; however, most existing studies have focused on metastatic disease or nephrectomy cohorts rather than SRMs or RMB specimens.38
This review has several limitations. The literature search was restricted to PubMed/MEDLINE and English-language publications, which may have excluded relevant studies indexed in other databases or published in other languages. Study selection and data extraction were performed by the authors without formal dual-reviewer screening, consistent with the narrative review methodology but introducing potential selection bias. The heterogeneity of study designs, patient populations, and endpoints across included studies precluded quantitative meta-analysis.
Conclusions
Molecular and genomic biomarkers have substantially advanced our understanding of RCC biology and highlight potential avenues to improve risk stratification for SRMs beyond conventional imaging and histopathologic assessment. Gene expression signatures such as ClearCode34 and the 16-gene recurrence assay, mutation-based markers including BAP1 and PBRM1 loss, chromosomal alterations such as 9p loss, and emerging liquid biopsy approaches collectively illustrate the promise of molecular characterization in refining SRM management. However, despite these advances, molecular biomarkers cannot currently be incorporated into routine clinical care for SRMs. Most candidate biomarkers have been developed in nephrectomy cohorts and have not been prospectively validated in RMB specimens or active surveillance populations. Consequently, their clinical utility for guiding surveillance or treatment decisions remains uncertain. Future progress will require prospective multicenter studies focused on SRM populations, standardized biopsy protocols optimized for molecular profiling, and validation of composite molecular risk models. Until such data are available, molecular biomarkers should be viewed as promising research tools rather than clinically actionable tests.
References
Siegel RL, Kratzer TB, Giaquinto AN, Sung H, Jemal A. Cancer statistics, 2025. CA Cancer J Clin. Jan–Feb 2025;75(1):10–45. doi:10.3322/caac.21871
Sanchez A, Feldman AS, Hakimi AA. Current Management of Small Renal Masses, Including Patient Selection, Renal Tumor Biopsy, Active Surveillance, and Thermal Ablation. J Clin Oncol. Dec 20 2018;36(36):3591–3600. doi:10.1200/jco.2018.79.2341
Hsieh JJ, Purdue MP, Signoretti S, et al. Renal cell carcinoma. Nat Rev Dis Primers. Mar 9 2017;3:17009. doi:10.1038/nrdp.2017.9
Welch HG, Skinner JS, Schroeck FR, Zhou W, Black WC. Regional Variation of Computed Tomographic Imaging in the United States and the Risk of Nephrectomy. JAMA Intern Med. Feb 1 2018;178(2):221–227. doi:10.1001/jamainternmed.2017.7508
Pierorazio PM, Johnson MH, Ball MW, et al. Five-year analysis of a multi-institutional prospective clinical trial of delayed intervention and surveillance for small renal masses: the DISSRM registry. European urology. 2015/09// 2015;68(3):408–415. doi:10.1016/j.eururo.2015.02.001
Bhindi B, Thompson RH, Lohse CM, et al. The Probability of Aggressive Versus Indolent Histology Based on Renal Tumor Size: Implications for Surveillance and Treatment. Eur Urol. Oct 2018;74(4):489–497. doi:10.1016/j.eururo.2018.06.003
Moch H, Amin MB, Berney DM, et al. The 2022 World Health Organization Classification of Tumours of the Urinary System and Male Genital Organs-Part A: Renal, Penile, and Testicular Tumours. Eur Urol. Nov 2022;82(5):458–468. doi:10.1016/j.eururo.2022.06.016
Frank I, Blute ML, Cheville JC, Lohse CM, Weaver AL, Zincke H. Solid renal tumors: an analysis of pathological features related to tumor size. J Urol. Dec 2003;170(6 Pt 1):2217–20. doi:10.1097/01.ju.0000095475.12515.5e
Campbell SC, Clark PE, Chang SS, Karam JA, Souter L, Uzzo RG. Renal Mass and Localized Renal Cancer: Evaluation, Management, and Follow-Up: AUA Guideline: Part I. J Urol. Aug 2021;206(2):199–208. doi:10.1097/ju.0000000000001911
Smaldone MC, Kutikov A, Egleston BL, et al. Small renal masses progressing to metastases under active surveillance: a systematic review and pooled analysis. Cancer. Feb 15 2012;118(4):997–1006. doi:10.1002/cncr.26369
Jewett MA, Mattar K, Basiuk J, et al. Active surveillance of small renal masses: progression patterns of early stage kidney cancer. Eur Urol. Jul 2011;60(1):39–44. doi:10.1016/j.eururo.2011.03.030
Richard PO, Jewett MA, Tanguay S, et al. Safety, reliability and accuracy of small renal tumour biopsies: results from a multi-institution registry. BJU Int. Apr 2017;119(4):543–549. doi:10.1111/bju.13630
Marconi L, Dabestani S, Lam TB, et al. Systematic Review and Meta-analysis of Diagnostic Accuracy of Percutaneous Renal Tumour Biopsy. Eur Urol. Apr 2016;69(4):660–673. doi:10.1016/j.eururo.2015.07.072
Brooks SA, Brannon AR, Parker JS, et al. ClearCode34: A prognostic risk predictor for localized clear cell renal cell carcinoma. Eur Urol. Jul 2014;66(1):77–84. doi:10.1016/j.eururo.2014.02.035
Rini B, Goddard A, Knezevic D, et al. A 16-gene assay to predict recurrence after surgery in localised renal cell carcinoma: development and validation studies. Lancet Oncol. Jun 2015;16(6):676–85. doi:10.1016/s1470-2045(15)70167-1
Morgan TM, Mehra R, Tiemeny P, et al. A Multigene Signature Based on Cell Cycle Proliferation Improves Prediction of Mortality Within 5 Yr of Radical Nephrectomy for Renal Cell Carcinoma. Eur Urol. May 2018;73(5):763–769. doi:10.1016/j.eururo.2017.12.002
Vaicekauskaitė I, Žalimas A, Sabaliauskaitė R, et al. Genomic analysis of small renal masses reveals mutations linked with renal cell carcinoma and fast-growing tumors. Journal of Cancer Research and Clinical Oncology. 2025/03/22 2025;151(3):118. doi:10.1007/s00432-025-06162-5
Comprehensive molecular characterization of clear cell renal cell carcinoma. Nature. Jul 4 2013;499(7456):43–9. doi:10.1038/nature12222
Sato Y, Yoshizato T, Shiraishi Y, et al. Integrated molecular analysis of clear-cell renal cell carcinoma. Nat Genet. Aug 2013;45(8):860–7. doi:10.1038/ng.2699
Gnarra JR, Tory K, Weng Y, et al. Mutations of the VHL tumour suppressor gene in renal carcinoma. Nat Genet. May 1994;7(1):85–90. doi:10.1038/ng0594-85
Kaelin WG, Jr. Molecular basis of the VHL hereditary cancer syndrome. Nat Rev Cancer. Sep 2002;2(9):673–82. doi:10.1038/nrc885
Ricketts CJ, De Cubas AA, Fan H, et al. The Cancer Genome Atlas Comprehensive Molecular Characterization of Renal Cell Carcinoma. Cell Rep. Apr 3 2018;23(1):313–326.e5. doi:10.1016/j.celrep.2018.03.075
Kapur P, Peña-Llopis S, Christie A, et al. Effects on survival of BAP1 and PBRM1 mutations in sporadic clear-cell renal-cell carcinoma: a retrospective analysis with independent validation. Lancet Oncol. Feb 2013;14(2):159–167. doi:10.1016/s1470-2045(12)70584-3
Peña-Llopis S, Vega-Rubín-de-Celis S, Liao A, et al. BAP1 loss defines a new class of renal cell carcinoma. Nat Genet. Jun 10 2012;44(7):751–9. doi:10.1038/ng.2323
Hakimi AA, Ostrovnaya I, Reva B, et al. Adverse outcomes in clear cell renal cell carcinoma with mutations of 3p21 epigenetic regulators BAP1 and SETD2: a report by MSKCC and the KIRC TCGA research network. Clin Cancer Res. Jun 15 2013;19(12):3259–67. doi:10.1158/1078-0432.Ccr-12-3886
Joseph RW, Kapur P, Serie DJ, et al. Clear Cell Renal Cell Carcinoma Subtypes Identified by BAP1 and PBRM1 Expression. J Urol. Jan 2016;195(1):180–7. doi:10.1016/j.juro.2015.07.113
Turajlic S, Xu H, Litchfield K, et al. Deterministic Evolutionary Trajectories Influence Primary Tumor Growth: TRACERx Renal. Cell. Apr 19 2018;173(3):595–610.e11. doi:10.1016/j.cell.2018.03.043
Turajlic S, Xu H, Litchfield K, et al. Tracking Cancer Evolution Reveals Constrained Routes to Metastases: TRACERx Renal. Cell. Apr 19 2018;173(3):581–594.e12. doi:10.1016/j.cell.2018.03.057
La Rochelle J, Klatte T, Dastane A, et al. Chromosome 9p deletions identify an aggressive phenotype of clear cell renal cell carcinoma. Cancer. Oct 15 2010;116(20):4696–702. doi:10.1002/cncr.25279
Brunelli M, Eccher A, Gobbo S, et al. Loss of chromosome 9p is an independent prognostic factor in patients with clear cell renal cell carcinoma. Mod Pathol. Jan 2008;21(1):1–6. doi:10.1038/modpathol.3800967
Monzon FA, Alvarez K, Peterson L, et al. Chromosome 14q loss defines a molecular subtype of clear-cell renal cell carcinoma associated with poor prognosis. Mod Pathol. Nov 2011;24(11):1470–9. doi:10.1038/modpathol.2011.107
Jani CT, Abdallah N, Tan A, Mckay RR. Liquid biopsy for renal cell carcinoma: A comprehensive review of techniques, applications, and future prospects. Kidney Cancer. 2024;8(1):205–225. doi:10.1177/24684570241303346
Pal SK, Sonpavde G, Agarwal N, et al. Evolution of Circulating Tumor DNA Profile from First-line to Subsequent Therapy in Metastatic Renal Cell Carcinoma. Eur Urol. Oct 2017;72(4):557–564. doi:10.1016/j.eururo.2017.03.046––
Peng YL, Yu B, Huang TX, et al. Early detection of renal cell carcinoma: a novel cell-free DNA fragmentomics-based liquid biopsy assay. ESMO Open. Jul 2025;10(7):105323. doi:10.1016/j.esmoop.2025.105323
Gerlinger M, Rowan AJ, Horswell S, et al. Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N Engl J Med. Mar 8 2012;366(10):883–892. doi:10.1056/NEJMoa1113205
Wan JCM, Massie C, Garcia-Corbacho J, et al. Liquid biopsies come of age: towards implementation of circulating tumour DNA. Nature Reviews Cancer. 2017/04/01 2017;17(4):223–238. doi:10.1038/nrc.2017.7
Ferro M, Musi G, Marchioni M, et al. Radiogenomics in Renal Cancer Management—Current Evidence and Future Prospects. International Journal of Molecular Sciences. 2023;24(5):4615.
Hamad RS, Sayed GA, Abd-Elmawla MA, et al. Targeting miRNAs in renal cell carcinoma: emerging therapeutic strategies. Int J Clin Oncol. Oct 2025;30(10):1925–1945. doi:10.1007/s10147-025-02856-5
Abbreviations
AUA — American Urological Association
AUC — area under the curve
CCP — cell cycle proliferation
ccRCC — clear cell renal cell carcinoma
cfDNA — cell-free DNA
ctDNA — circulating tumor DNA
EAU — European Association of Urology
IHC — immunohistochemistry
MeSH — Medical Subject Headings
RCC — renal cell carcinoma
RMB — renal mass biopsy
SRM(s) — small renal mass(es)
TCGA — The Cancer Genome Atlas
Lorem ipsum dolor sit amet, consectetur adipiscing elit. Ut elit tellus, luctus nec ullamcorper mattis, pulvinar dapibus leo.