Submitted - AUGUST 2, 2023 | Revised AUGUST 29, 2023 Accepted -AUGUST 30, 2023 | ePublished - OCTOBER 10, 2022
https://doi.org/10.52733/KCJ21n3-r2

ABSTRACT
Human endogenous retroviruses (hERVs) have emerged as a mechanism for tumor development and progression in clear cell renal cell carcinoma (ccRCC). Increased expression of various hERVs has been reported in ccRCC with associated activation of anti-tumor immune responses. Retrospective analysis of hERV expression in human ccRCC tumor tissue suggests hERV expression may be associated with improved response to immune checkpoint inhibitors. However, the use of expression to predict response is limited by our ability to annotate and detect hERV expression. This review discusses the biology of hERVs, their role in ccRCC, and the possible impact on ccRCC response to immunotherapy.
KEYWORDS
Renal Cell Carcinoma, Endogenous Retroviruses, Immunotherapy
INTRODUCTION
Kidney cancer is the eighth most
common cancer among both
sexes in the United States and
is estimated to cause 14,890 deaths
in 20231. Clear cell renal cell carcinoma
(ccRCC) is the most common
histologic type of kidney cancer,
comprising up to 85% of RCC.
ccRCC is characterized by the loss or
mutation of the von Hippel-Lindau
gene, resulting in constitutive activation
of hypoxia-inducible factors
(HIF) and upregulation of downstream
signaling pathways, including
vascular endothelial growth factor
(VEGF). Other commonly mutated
genes in ccRCC include those that
encode chromatin-modifying enzymes,
such as SETD2, PBRM1, and
BAP-1, and PIK3CA. Over the past
20 years, the treatment paradigm
for ccRCC has substantially changed
with improved understanding of the
underlying tumor biology. However,
a mainstay in systemic therapies for
ccRCC has been immunotherapy
with a relative lack of understanding
of the biologic drivers of response
and resistance in ccRCC.
Historically, ccRCC has been considered responsive to immunotherapy with interferonalfa and high-dose interleukin-2 as standard treatments 2,3 . More recently, ccRCC has demonstrated significant response to immune checkpoint inhibitors (ICI), but activity is only observed in a subset of tumors. A proposed mechanism of ICI response in other tumors is high tumor mutational burden (TMB) leading to increased tumorassociated antigens. In melanoma, increased TMB is associated with significantly improved longterm benefit 4. However, ccRCC demonstrates a lower TMB than other cancers that respond to ICI. For example, melanoma typically has 10- 400 mutations per megabase4, while ccRCC demonstrates an average of 1.1 mutations / Mb 5-7. Since ccRCC has lower TMB, alternative mechanisms of immunogenicity have been evaluated and expression of human endogenous retroviruses (hERVs) have been identified as a possible biomarker of response. Over the past couple of decades, hERVs have been increasingly recognized as upregulated in human cancers 8-16. Additionally, hERV products have been shown to elicit antitumor immune response in both renal cell carcinoma and other tumor types 17-22. Recent studies highlight the significant role that hERVs may play not only in the development and progression of ccRCC, but also the response to immunotherapy 15,23–25. In this review, we focus on the biology of hERVs, their identified roles in RCC, and how hERVs may impact response to immunotherapy in RCC.
The biology of endogenous retroviruses
Human endogenous retroviruses (hERVs) are endogenous viral components present in the human genome which originated as retroviruses millions of years ago and were incorporated into the genome of germ line cells. hERVs form the majority of long terminal repeats (LTRs) and comprise about 8% of the human genome 26. While hERVs are defective in viral replication and typically lose the ability to encode proteins, they contribute to regulation of the human genome by acting as promoters, enhancers, repressors, poly-A signals, and alternative splice sites for human genes19. hERVs are typically silenced in normal somatic tissues19, but hERV expression has been reported as increased in a variety of cancers 8–14, including ccRCC 15,17,18, autoimmune disease, and neurological disorders 27-30.
Over 50 families of hERVs
have been identified and are
categorized into classes I-III 26. For
example, HERV-E and HERV-H
are class I, while HERV-K is a class
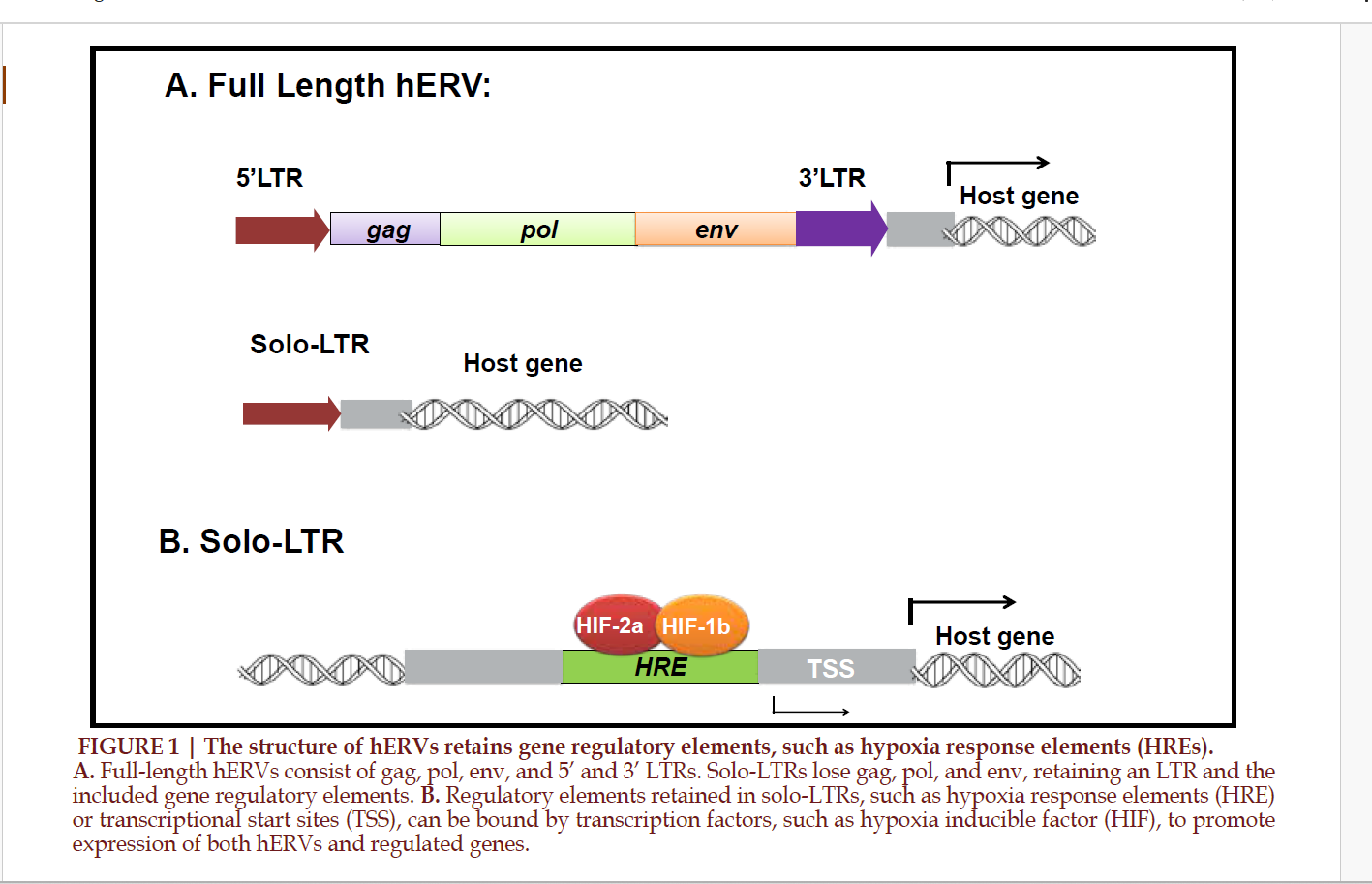
II hERV26. The structure of each
individual hERV typically contains
gag, pol, and env components, which
are flanked on the 5’ and 3’ ends
by two gene regulatory sequences,
long terminal repeats (LTR) 26.
While most of the hERVs in the
human genome lose coding ability,
a few hERVs retain the ability to
encode functional proteins, such
as HERV-K and HERV-W 31,32.
Loss of hERV coding ability can
be due to non-allelic homologous
recombination between the 3’ and
5’ LTRs, resulting in solo-LTRs
and loss of the gag, pol, and env
components 33,34. Within the human
genome, hERVs typically exist in
the solo-LTR form and maintain
gene regulatory function through
the presence of transcriptional
regulatory motifs 34,35 (FIGURE 1).
However, some hERVs, such as those
in the ERVK family, do preserve a
functional gag gene or open-reading
frame for the pol and env genes 36.
hERVs may promote tumorigenesis through a variety of mechanisms. First, expression of hERVs can activate tumor-promoting signaling pathways, including the RAS-ERK and Wnt/β-catenin pathways 8,37,38, which promote cell proliferation and transformation. Second, the hERV envelope protein, syncytin-2, has been shown to have immunosuppressive properties 39. However, hERV expression also promotes the detection of tumors by the immune system. Immunotherapy research in other tumor types has demonstrated that a subset of HERV-K and HERV-H proviruses express immunestimulating antigens on tumor cells, which can then be recognized and killed by cytotoxic T-cells 20,22.
Endogenous retroviruses in clear cell renal cell carcinoma
Over the past two decades, hERV expression has been strongly implicated in the development and progression of ccRCC and is associated with clinical outcomes. First, multiple hERVs demonstrate increased expression in ccRCC, including HERV-E 16,18, HHLA2 40, and HERVERI 41. Interestingly, expression of HERV-E in ccRCC appears to be interrelated to the underlying tumor biology. HERV-E expression levels correlate with HIF- 2α levels and HERV-E expression was abrogated by introduction of normal VHL or HIF-2α knockdown16. Additionally, HIF-2α can act as a transcriptional factor for HERV-E by binding a HIF response element (HRE) located in the proviral 5’ long terminal repeat (LTR) 16. Cherkasova et al., also demonstrated that this LTR was hypermethylated in normal tissues, preventing hERV expression, and hypomethylated in HERV-E expressing ccRCC tumors 16, allowing for increased expression. In a separate study, Siebenthall et al identified HIF-binding to other LTR sites genome-wide which correlated with gene expression changes in RCC, including HIF binding at an HRE in an hERV LTR located upstream of the stem cell transcription factor POU5F1 (OCT4), resulting in increased POU5F1 expression levels 42.
Increased hERV expression
is also associated with PBRM1 loss
in primary human ccRCC tumors41.
PBRM1 is the second most frequently
mutated gene in ccRCC5 and encodes
a member of the PBAF (polybromo
BRG1 associated factor) SWI/SNF
chromatin remodeling complex 43,44.
This SWI/SNF complex regulates
nucleosome positioning and gene
expression 43,44. We utilized the
UMRC2 kidney cancer cell line to
confirm that in vitro silencing of
PBRM1, HIF1, and HIF2 resulted in
increased expression of hERVs in a
HIF1 a nd HI F2 dependent manner41.
We also identified a specific family of
hERVs, the HERVER I superfamily,
that are enriched in PBR M1-regulated
hERVs 41. Therefore, expression of the
HERVERI super family is dependent
upon loss of function mutations in
two genes that a re highly specific to
ccRCC, VHL a nd PBRM1, and may
explain its unique association with
this cancer.
Furthermore, the expression
of hERVs in ccRCC is immunogenic,
activating T-cell responses. First,
in a study utilizing TCGA datase ts
f rom 18 tumor types, Rooney et
al. identified that hig h immune
cytolytic activity in ccRCC is
associated with elevated expression
of the HERV-E loci, ERVE-4 45.
Additionally, Cherkasova et al.
demonstrated that proteins predicted
to encode the HERV-E envelope
protein (HLA-A*0201-restricted
peptides) are expressed in ccRCC
tumors and are immunogenic in
vitro 17. Furthermore, in a patient
demonstrating regression of renal
cell carcinoma after receiving an
allogeneic hematopoietic stem cell
transplant, a CD8+ T-cell clone
recognizing a HERV-E antigen was
isolated 18, suggesting tumor-specific
T-cell reactivity in response to
HERV-E expression. These results
indicate that hERV- based antigens
could act as targets for possible
T-cell derived immunotherapy in
ccRCC.
Finally, the expression of
hERVs in ccRCC is associated with
patient clinical outcomes. Human
endogenous retrovirus-H long
terminal repeat-associating protein
2 (HHLA2) demonstrates increased
expression in ccRCC compared to
normal kidney tissue at both RNA
and protein levels 40 and HHLA2
expression was associated with poor
overall survival 40. Additionally, in
a study utilizing the TCGA (The
Cancer Genome Atlas) pan-cancer
dataset, mean hERV expression in
ccRCC was significantly negatively
prognostic for overall survival
and, when comparing Kaplan
Meier curves for the upper versus
lower 50th percentile mean hERV
expression, ccRCC was one of only
five tumor types that demonstrated
significant separation of survival
curves 15. Of these five tumor
types, ccRCC demonstrated the
most significant association, with
higher hERV expression associated
with significantly shorter overall
survival15. Further work in this
dataset identified possible hERV
signaling through the RIG-I-like
pathway and B-cell activation and
patients with both higher expression
of B-cell receptor-associated
signatures and down-regulation of
RIG-I-like signatures demonstrated
significantly shorter overall
survival15.
The impact of ERVs on response to immunotherapy in RCC
The introduction of immune checkpoint inhibitors (ICI) for the treatment of ccRCC has significantly improved patient outcomes. However, significant responses are only observed in a subset of patients and much work has focused on identifying predictive biomarkers. Given the immunogenicity of hERV expression discussed above, studies have utilized patient samples from ICI clinical trials to assess the association between hERV expression and tumor response to ICI.
In 24 metastatic ccRCC
tumors treated with singleagent
PD-1/PD-L1 blockade,
ICI responders demonstrated
significantly higher expression of
ERV3-2 than non-responders23.
Using the TCGA KIRC dataset,
this study also demonstrated that
high expression of twenty hERVs
that were identified as potentially
immunogenic was associated with
increased immune infiltration,
checkpoint pathway upregulation,
and a higher CD8+ T-cell proportion
in tumor infiltrating leukocytes
compared to low hERV expression23.
By performing qRT-PCR on tumor
samples from CheckMate010,
Pignon et al. also evaluated the
association between 4 hERVs (pan-
ERVE4, pan-ERV3.2, hERV4700
GAG, and hERV4700 ENV) and
response to nivolumab 24. Using a
cutoff of the 25th percentile, high
levels of hERV4700 ENV were
associated with significantly longer
median progression free survival
and higher overall response rates24.
Similarly, using tumor samples
from CheckMate 025, Ficial et al.
identified that in ccRCC tumors
treated with nivolumab, higher
hERV-E RNA expression levels were
associated with increased durable
response rate and longer progressionfree
survival25. Additionally, in the
previously mentioned TCGA pancancer
dataset, a transcriptional
signature indicating anti-PD1
responsiveness (IPRES_aPD1_
responder) demonstrated positive
association with hERV expression
in 79.2% of significantly associated
hERVs in all tumor types15. Within
ccRCC specifically, higher expression
of hERV 4700 was associated with
response to anti-PD1 therapy 15.
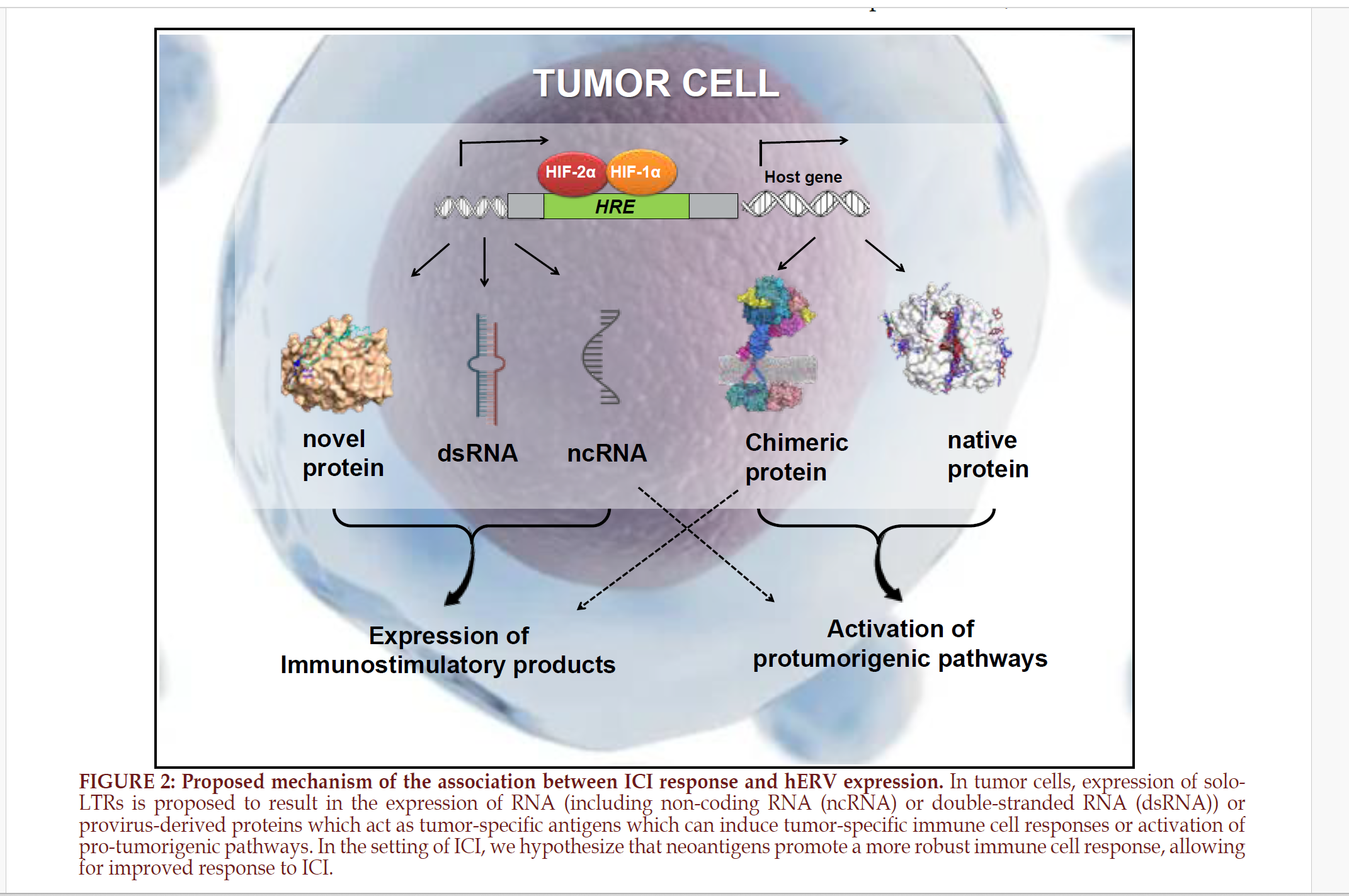
When combined, these studies
suggest that high hERV expression
may identify patients who might
respond to ICI. FIGURE 2 illustrates
a proposed mechanism for this
improved response in the setting of
hERV expression.
However, when Braun et
al., subsequently pooled data from
CheckMate009, CheckMate010, and
CheckMate025, they did not identify
a robust association between
hERV expression and response
to immunotherapy. In this study,
they first validated RNA-seq-based
expression of hERV using qRTPCR
and demonstrated that RNAsequencing
did not reliably quantify
ERV3-2 expression. However, they
did identify a weak association
between ERV2282 and ERV3382
expression with response and
overall survival and progression free
survival. However, when divided
into high and low expression levels,
the significant association with PFS
and OS did not persist46.
Additionally, using tissue
from the ADAPTeR trial, in which
patients with metastatic ccRCC
were treated with nivolumab, Au
et al concluded that ccRCC-specific
hERV expression did not directly
correlate with response to anti-
PD-1 treatment 47. Specifically, they
performed RNA-sequencing on a
total of 60 tumor samples from 14
patients and annotated hERVs using
a previously built “complete custom
repeat region annotation” 48. Even
when accounting for annotation
discrepancies between prior
analyses, the hERVs previously
identified as associated with
cytotoxic T-cell presence, ccRCC
response to ICI, or providing antigens
were not differentially expressed
between ICI responders and nonresponders
or associated with ICI
response in this study 47. However,
10 different hERV annotations were
significantly associated with ICI
response but demonstrated a mix
of restriction to responders versus
non-responders, demonstrating a
different pattern of hERV association
with ICI response than observed
in the above studies 47. Based on
these results and data indicating
that hERVs previously reported
as upregulated in ccRCC may be
expressed on immune cells, Au et
al suggest that hERV expression in
ccRCC may reflect tumor purity and
the diverse cellular composition of
ccRCC tumors 47.
As described above,
PBRM1 loss is associated with
increased expression of hERVs in
primary ccRCC human tumors
and additional work has evaluated
the interplay between PBRM1
mutation, hERV expression, and ICI
response. First, previous work has
evaluated predictors of ICI response
in ccRCC and variably identified
PBRM1 mutations as a predictive
biomarker 46,49–53. While studies
identified an association between
PBRM1 loss of function mutations
and second-line, single-agent ICI
response 46,49,50,53, additional groups
evaluating PBRM1 mutations
and ICI response in first-line
treatment with combination VEGF
inhibitor and ICI did not identify an
association 51,52. Additional work by
Liu et al highlights the role that HIF
plays in this response since PBRM1
deficient, HIF axis-intact cells show
ICI resistance 54. This study utilized
VHL and PBRM1 wild-type RENCA
cells, which are murine-derived RCC
cells from a BALB/c background,
in which PBRM1 knockout was
achieved using CRISPR/Cas9
technology 54. When introduced
into mice subcutaneously, both
PBRM1 wild-type and knockout
cells established tumors and
PBRM1 knockout tumors showed
worse survival than control tumors
following treatment with PD-1
antibody 54. Further evaluation of
how the concurrent loss of PBRM1
and VHL impact ICI response is
needed.
In addition to using hERV
expression as a predictive biomarker
for ICI response, future directions
can also explore alternative
approaches to exploiting the biology
of hERVs. First, as hERVs are
immunogenic, they may have the
capacity to serve as vaccine targets.
Indeed, in a mouse model with
tumors formed from murine renal
carcinoma cells (Renca) altered to
express the HERV-K Gag proteins,
mice vaccinated using a recombinant
virus expressing the HERV-K Gag
protein demonstrated reduced
tumor growth and reduction in
pulmonary tumor nodules55. Similar
results were observed when mice
with tumors expressing HERV-K
Env proteins were vaccinated
against the HERV-K Env protein 56.
Second, it may also be possible
to manipulate the expression of
hERVs to increase response to
immunotherapy. For example,
kidney cancer cell lines and primary
cells that were treated with a DNA
hypomethylating agent, decitabine,
demonstrated increased expression
of transposable elements, LINE1,
and ERVs ERV3-2 and ERV4700,
which were associated with immune
infiltration and ICI response on
bioinformatic analysis 57. Finally,
work investigating the impact of
treating HLA-A*11:01 positive
patients with metastatic ccRCC with
HERV-E TCR transduced CD8+ and
CD34+ enriched T-cells is ongoing
(NCT03354390) and remains a
promising option for exploiting
hERV expression to more effectively
treat ccRCC.
CONCLUSIONS
A subset of ccRCC tumors demonstrate increased expression of human endogenous retroviruses, endogenous viral components which have been incorporated into the human genome. ccRCC expression of hERVs seems to be interrelated to its distinct underlying tumor biology, with hERV expression levels related to both the VHLHIF pathway and PBRM1 loss. Furthermore, the expression of hERVs in ccRCC is immunogenic, resulting in activation of tumorspecific T-cell responses in vitro and in vivo, and studies in mouse models highlight the potential for hERVs to act as vaccine targets. While higher hERV expression is associated with worse overall survival in ccRCC, data evaluating the association between hERV expression and response to ICI is conflicting. While single study reports identified encouraging associations with improved patient outcomes, only weak associations were observed when studies were combined, possibly reflecting differences in intratumoral heterogeneity and the tumor microenvironment. As such, additional knowledge of the mechanisms and pathways by which HERVs impact ccRCC tumorigenesis and therapeutic response is needed for optimal therapeutic development and continued improvements in patient outcomes.
FUTURE DIRECTIONS
Further investigation of the impact of human ERVs on the pathogenesis and progression of ccRCC will allow for improved understanding of the role ERVs play in response to therapies. Additionally, utilizing tissue from clinical trials assessing response to combination immunotherapy or prior to receiving systemic therapy may shed light on the seeming discrepancies in the association of hERV expression and ICI response. Finally, a broader understanding of the biology of hERV in ccRCC is necessary, including 1) characterizing the expression of hERVs in ccRCC tumor cells versus the tumor microenvironment; 2) elucidating the key downstream signaling pathways activated by hERVs and the interplay with VHL loss and chromatin modifying enzymes, and 3) identifying additional tumor-specific antigens. Further knowledge of the key cell types, antigens, and signaling pathways impacted by hERVs will allow further development of synergistic therapies and optimization of first-line treatments for individual patients.
FUNDING STATEMENT
This work was supported by funding from the National Institutes of Health (UNC Integrated Translational Oncology Program T32-CA244125 to UNC/khg). TLR is supported by the National Cancer Institute at the National Institutes of Health (grant number 1K08CA248967-01).
REFERENCE
# Corresponding Author: Marc A Bjurlin in ccRCC. Department of Urology, University of North Carolina at Chapel Hill, Chapel Hill, NC. Email: marc_bjurlin@med.unc.edu